Research Overview
The extracellular matrix (ECM), a complex meshwork of cross-linked proteins, is a fundamental component of multicellular organisms. It provides architectural support to cells, confers mechanical properties to tissues, and conveys biochemical signals transduced by cell-surface receptors to control various cellular processes such as proliferation, survival, differentiation, adhesion, and migration. Alterations in the composition and organization of the ECM cause or accompany the development of diseases such as fibrosis, cardiovascular diseases, and cancer (Naba, Nat Rev Mol Cell Biol, 2024).
To explore the importance of the ECM in diseases, our lab has developed transformative proteomic and bioinformatic approaches to study the molecular composition of the ECM, also known as the “matrisome,” in normal and diseased tissues.
Using these approaches, we discovered that the ECM of any given tissue is composed of 150+ proteins, and that ECM composition varies between tissues, between normal tissues and tumors arising from them, between tumors of different metastatic potential, and between primary tumors and their distant metastases. Importantly, our pipeline is a powerful tool for devising hypothesis-driven research, as it has led to the discovery of a novel ECM protein called SNED1 (Naba et al., eLife, 2014).
Our current research centers around three themes:
Development of novel proteomic and bioinformatic methods for deep matrisome profiling
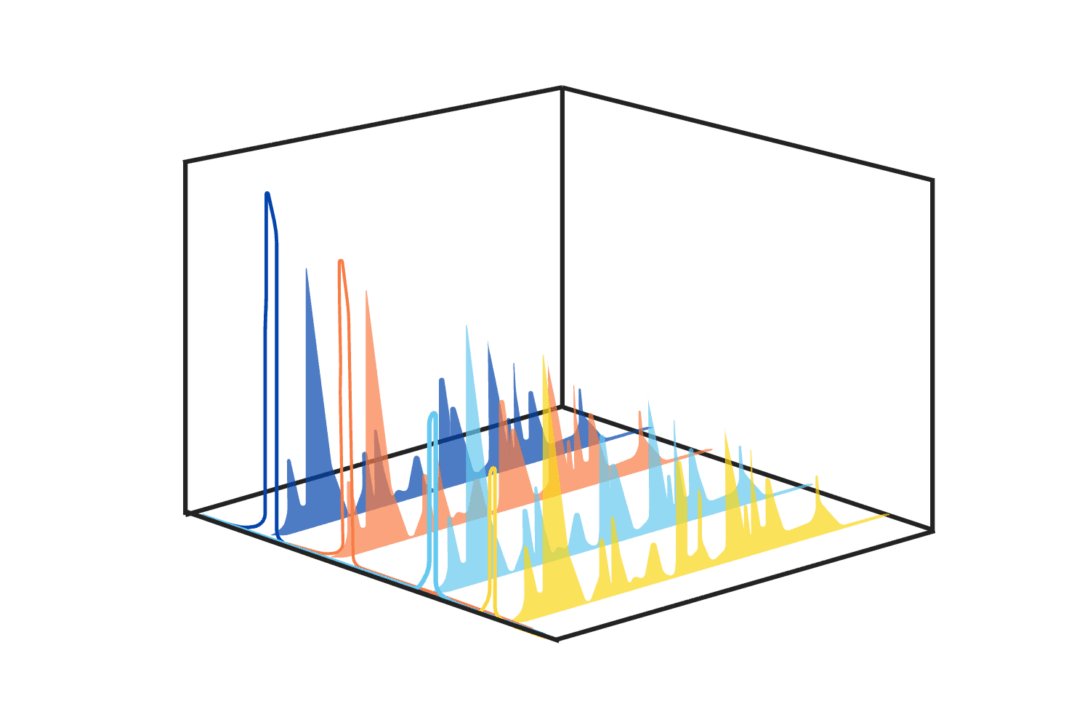
While our work has contributed to making proteomics a state-of-the-art approach to study the global composition of the ECM of tissues (Naba, Mol Cell Prot, 2023), many critical aspects of the ECM remain to be discovered, including the nature of ECM protein proteoforms and post-translational modifications of the protein interactions occurring in different pathophysiological contexts.
To address these remaining gaps, we are continuing to develop novel methods to achieve what we term “deep matrisome profiling”.
Visit the Matrisome Project website to learn more.
Recent Publications
- Leverton L, Bains AK, Isa I, Ricard-Blum S, Naba A. A guide to building the matrisome interactome: From computational prediction to experimental validation. FEBS Journal, 2025, 293(1):42-70. Journal Access | PMC Access
- Lamba R, Paguntalan AM, Petrov PB, Naba A*, Izzi V*. MatriCom: a scRNA-Seq data mining tool to infer cell-extracellular matrix interactions. Journal of Cell Science, 2025, 138(13):jcs263927. Journal Access | PMC Access
- Bains AK and Naba A. Proteomic insights into the extracellular matrix: a focus on proteoforms and their implications in health and disease. Expert Review of Proteomics, 2024, 21(11):463–481. Journal Access | PMC Access
Relevant Funding
- NIH Human Biomolecular Atlas Program U01HG012680: Thinking outside the cell: Leveraging HuBMAP data to build the human ECM atlas
- NIH/NCI R21CA261642: Enhanced mass-spectrometry-based approaches for in-depth profiling of the cancer extracellular matrix
Exploring and exploiting the ECM in cancer
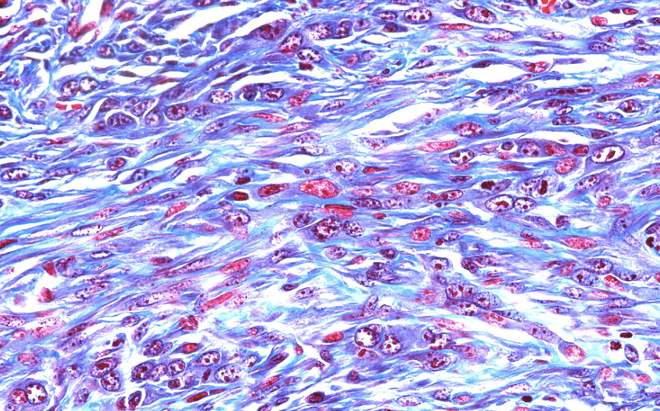
Using pre-clinical models and patient samples, our work has contributed to showing that the ECM plays key roles at every step of tumor progression, including tumor growth, the angiogenic switch, metastatic dissemination, and response to immunotherapy (Socovich and Naba, Semin Cell Dev Biol, 2018).
We are now interested in translating our findings to develop matritherapies and improve cancer patient outcomes.
Recent Publications
- Considine JM, Pally D, O’Brien SA, Potts, J, Feng D, Pignatelli J, Kashyap AS, Sharma N, Naba A. In-depth proteomic profiling of the extracellular matrix of pancreatic ductal adenocarcinomas identifies signatures correlating with lymphocyte infiltration. bioRxiv, 2025. bioRxiv Access
- Considine JM, Gomez C, Pally D, Yang N, Taha IN, Sorenson J, Brooks EG, Kreeger PK*, Naba A*. Comparative proteomic analysis of the ECM composition of the human omentum and mesentery, the main sites of ovarian cancer metastasis. iScience, 2025, 29(1):114461. Journal Access | PMC Access
- Di Martino J, Nobre AR, Mondal C, Taha IN, Farias E, Fertig E, Naba A, Aguirre-Ghiso J, and Bravo-Cordero JJ. A tumor-derived type III collagen-rich ECM niche regulates tumor cell dormancy. Nature Cancer, 2021, 3(1):90-107. Journal Access | PMC Access
Relevant Funding
- NIH/NCI R01CA290693 – Collaboration with Dr. Pam Kreeger (UW-Madison): Elevated collagen I and fibronectin in the ovarian cancer pre-metastatic niche
- Sponsored research project with Boehringer Ingelheim: Proteomic characterization of the changes in tumor ECM composition in response to anti-cancer therapies
Deciphering the roles of the novel ECM protein, SNED1, at the intersection of development and cancer metastasis
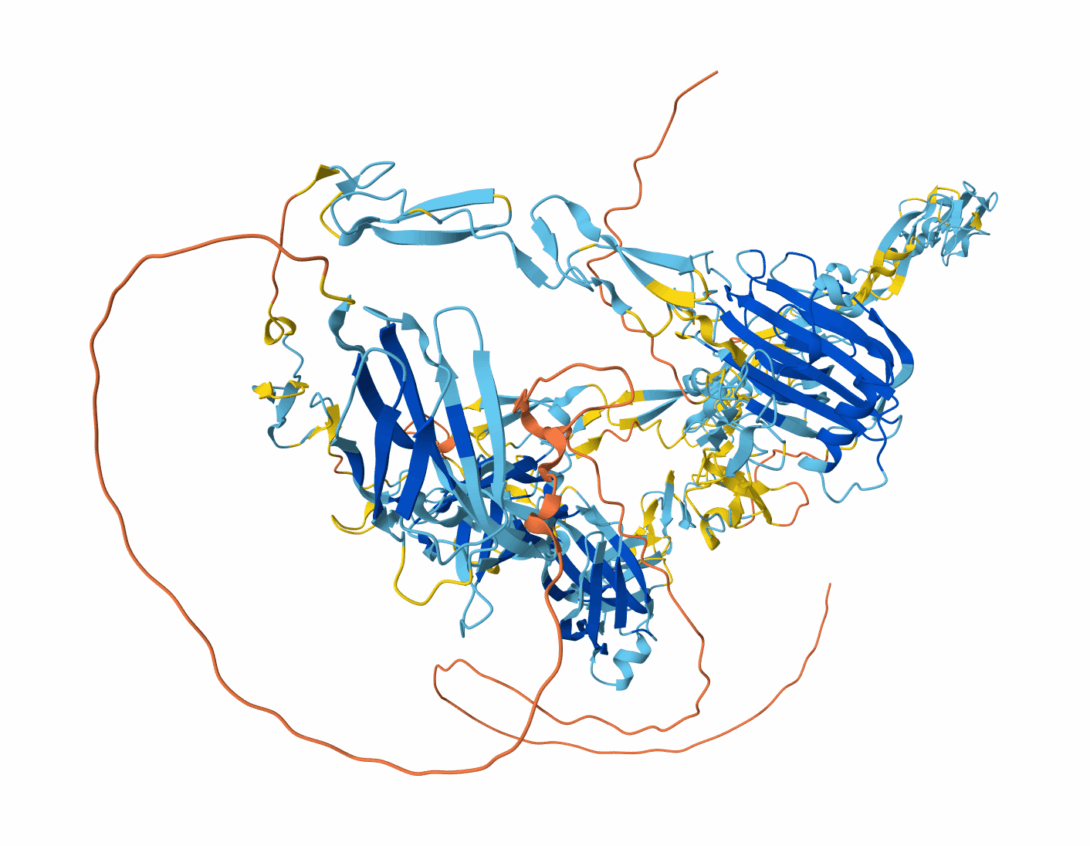
We previously identified a novel ECM protein, SNED1 (Sushi, Nidogen and EGF-like domain protein 1), in a proteomic screen aimed at identifying ECM proteins differentially expressed between highly and poorly metastatic mammary tumors. We found that SNED1 played a functional role in tumor dissemination, as SNED1 knockdown decreased metastasis in a mouse model of breast cancer. We further showed that the level of expression of SNED1 was a prognostic factor for hormone receptor-negative breast cancer patients (Naba et al., eLife, 2014). Our goals are now to identify the mechanisms controlled by SNED1 contributing to breast cancer metastasis and to evaluate the prognostic value of SNED1 expression for human cancer patients.
The importance of SNED1 in breast cancer metastasis prompted us to investigate its roles in development and physiology. To do so, we generated the first knockout mouse model of Sned1 and demonstrated that Sned1 is an essential gene, as its deletion resulted in early neonatal lethality due to severe craniofacial malformations. Our current work focuses on deciphering the mechanisms of SNED1 assembly in the ECM and the impact of SNED1 on cellular processes, like epithelial-to-mesenchymal transition (EMT) and migration, involved in craniofacial morphogenesis.
Recent Publications
- Pally D, Kapoor N, Naba A. The novel ECM protein SNED1 mediates cell adhesion via the RGD-binding integrins α5β1 and αvβ3. Journal of Cell Science, 2025, 138(2):JCS263479. Journal Access | PMC Access
- Vallet SD, Davis MN, Barqué A, Thahab AH, Ricard-Blum S, and Naba A. Computational and experimental characterization of the novel ECM glycoprotein SNED1 and prediction of its interactome. Biochemical Journal, 2021, 478(7):1413–1434. Journal Access
- Barqué A, Jan K, De La Fuente E, Nicholas CL, and Hynes RO*, and Naba A*. Knockout of the gene encoding the extracellular matrix protein Sned1 results in craniofacial malformations and early neonatal lethality. Developmental Dynamics, 2020, 250(2):274-294. Journal Access | PMC Access
Relevant Funding
- NIH/NIGMS R01GM148423: Mechanisms guiding the fibrillar assembly of SNED1 in the extracellular matrix
- UIC Chancellor Translational Research Initiative: Targeting the interaction of the ECM protein SNED1 with its receptor to prevent breast cancer metastasis